"What is real? How do you define real? If you're talking about your senses, that you feel, taste, smell, or see, then all you're talking about are electrical signals interpreted by your brain." - Morpheus, The Matrix, 1999.
Monday, May 28, 2007
...phew !
a sense of belonging...what one does now is no longer going to be just personal curiosity...a concerted, directed effort backed by dedicated personnel and resources.
no trivial pursuits...not that they were ever so to oneself...but the way the world sees it...
one has now been absolved of the guilty pangs accompanying post-Lewin et al sessions...
i had coffee twice today. one frappe at the bangalore airport. one hot coffee aboard the Mumbai bound flight...
ones cAMP must be on overdrive now...caffeine can drive one crazy...phew !
Wednesday, May 16, 2007
Something On Genome Wide Repeats

Was readin up on chromosome organisation in Watson. Something came to my mind. Two things actually.
Transposon Dating: How rare are human transposition events? Can those sequences, which are relics of transposition events found in the genome-wide-repeat subset of the intergenetic DNA, be used for phylogenetic analysis ?
Gene Pool Hypothesis: DNA sequences in the genome wide repeat subset of the intergenetic zone are thought to be relics of transposition events. They play no functional role in the species with respect to lending their DNA as an expression template. Can one expect the evolutionary pressures to be relaxed on these zones relative to actual protein coding/regulatory genetic zones? If so, then could this rich pool of variable DNA sequences serve as a birth place of novel genes. A genetic nebula. Such novel genes, if they exist at all and are discovered would be expected to bear little homology to other pre-existing genes.
If a tree based on sequence homology for all the 27000 human genes is constructed what would it look like?
1.What is the kind of homology amongst these 27000 genes?
2.Out of them how many would have a true common single ancestral gene and how many would have formed independently of this ancestral root? Or how many roots can account for the entire human gene repertoire?
3.Would these independently formed nodes in this tree be good places to look for genes which makes humans more so than say other primates?
4.Have any of the above independently formed genes been derived from mutation in the transposition-events relics found in the genome-wide-repeat zones in the intergenetic DNA (discussed previously) ?
5.Can the above mentioned zones give rise to sequences coding for trans-regulatory-RNA apart from or if not full-fledged genes?
If any of the above 2 mechanisms are real then some of the complexity of humans can be ascribed to the above gene forming pools of the junk DNA component of the human genome.
Friday, December 01, 2006
Sun Java Application Server - MySQL : Connection Pooling HOWTO
This time it is...
MySQL Connection Pooling Step-by-Step HOWTO
- Make sure U have the mysql.jar file with U. It contains the mysql driver.
- Your Sun Java Application Server (SJAS) folder looks like this
- Now copy the mysql.jar to J2EE_HOME\ant\lib directory
- Start the server
- Open the admin consolehttp://localhost:4848/
- Login using the default uname/passwd
uname: admin
passwd: adminadmin
- Click on : Application Server => JVM Settings tab => Path Settings tab
- Now paste the following line: ${com.sun.aas.antLib}/mysql.jar in the textarea labeled as "Classpath Suffix"
- Click on Save. Close the admin console.
- Restart the server.
- Open the admin console (http://localhost:4848/) again.
- Expand the Resources node
- Expand the JDBC sub-node
- Click on Connection Pools
- Click on New
- Fill up the fields as follows:
Name: mysqlpool
Datasource Classname: com.mysql.jdbc.jdbc2.optional.MysqlDataSource
Resource Type: javax.sql.DataSource
Do not modify rest of the fields.
- Now add following properties by clicking on "Add Property" at the bottom of the form:
DatabaseName : OnlineBankingSystem
User : admin (your mysql username)
Password : admin (your mysql password)
- Click on save. You will see "New values successfully saved"
- Now, Click on Ping. If you get a "Ping succeeded." then be happy :D
- Now is the last step. click on the JDBC Resources node.
- Click on New.
- Fill up the fields as follows:
JNDI Name: mysql
Pool Name: mysqlpool
- Click on Save. Thats it! We r done ! :D
- Your mysql connection pool is now ready to accept requests to the database. It is referred to as just "mysql" from within the EJB and JSP (its JNDI Name remember, :-? ).
- And of course, I wish U all the Best ! :D
Lemme know soon wat happened and if it helped. :D
Sunday, September 03, 2006
What does a CD mean to a cow?

Passing thought: Imagine two people conversing using their Nokias. The information is ferried between them through space in form of light waves. Would it be possible to monitor the physical properties of the space and infer the information being exchanged? One may point out that the cell phone data is garbled for obvious reasons. But would it be possible to determine atleast this garbled data by performing a suitable measurement on the space, somehow?
Eavesdropping on people's conversations is not really the topic. It is rather about the nature of information.
Is there a theoretical limit to how much information a given volume of space may possibly
contain irrespective of the method used for storing/encoding this information? There can be two possibilies:
- Yes, there is a limit.
- No, there isn't a limit.
But if it happens that the second case is true then information would turn out to be indeed something very personal - so that each individual can have the freedom of interpreting this information the way way he pleases. That there wont be such thing as a universally true information or a universally false information for a given obeject space under scrutiny. In such a case it would not be possible to develop a generic tool or quantitating the information content Which is :-( So, clearly in this case infomation exchange is possible only if the two parties agree on the information content.
Passing thought: Your Win-Zip program which you use to compress data does its thing like this[2]. A CD contains data stored along the tracks. In each track there are two possible features - a small pit and no pit corressponding 0 and 1 or otherwise. A 700Mb of such a CD would contain an entire DivX of The Da vinci Code. One would be able to watch it if one has a CD-ROM to decode the engraved data. But what if one does not have such a specific device? Does the information contained in this CD affect the properties of the physical space it occupies so that a suitable measurement on this space would reveal the movie data indirectly?
Casual Mention: "Biology is the domain of the complex. It takes 3 * 10.pow(9) bases = 6 * 10.pow(9) bits of information to specify the DNA that determines a human being".[1]
But what is in it for the biologist? Well, putting it rather (very) bluntly,
If the information is found to be universal as opposed to personal then a given space, say a space which contains only a single DNA fragment, can be analysed by using some tool to come up with the absolute information content of the space; DNA sequence in this case. Outrageous. I know.
Theres more actually, but by now I have either forgotten to mention those things/ too lazy to type anymore/ its lunch time :P
Maybe some other day. . .
Interesting reads:
- http://www.cs.auckland.ac.nz/CDMTCS/chaitin/***
- http://en.wikipedia.org/wiki/Data_compression
- http://www.faqs.org/faqs/compression-faq/part2/section-4.html
*** = Must read.
Wednesday, June 07, 2006
T.I.F.R. BioChem Labs

Plush, clean, efficient publishing machines, silent except for the ho-humming of speed-vac rotors or gel-shakers amidst over-crowded chemical shelves with a graffiti of TODO post-its whereas the room corners are claimed by the latest and greatest fat gizmos spitting out raw data only to be handed over to the computing room to make sense of all that alphabet soup while the show is run by smart biochemical ensembles of CHONSP.
ΔG = ?
Tuesday, January 17, 2006
Million Dollar Meccano

Some Basics
Everybody knows the central dogma of molecular biology - DNA=>RNA=>Protein. There is another lesser know dogma - protein primary sequence=>protein 3D structure=>protein function. The real challenge now is to actually be able to predict a protein's function given its amino acid sequence.
Proteins come in mind boggling range of shapes and functions. Yet all of these exotic stuctures derive their blueprints from a modest (boring), 1D stretch of information encoded in a mRNA fragment. Hence, essentially, manufacturing a protein is simply (!) the folding up of a linear polymer of amino acids onto itself through twists and turns and coils assuming a complex 3D structure which can do useful things.
There is considerable evidence suggesting that the protein's primary structure contains sufficient information to direct the folding of the polypeptide backbone to give a correctly folded 3D sturcture.
Show Me My Millions
Biology never had it so good. We now seem to know much of the basics, we seem to have the suitable technology and most importantly we also have brains working towards this massive goal. It is just a matter of who claims the top cherry. Biology is therefore going to be an increasingly interesting area even for the VCs. The X Pize Foundation of the first-private-spaceship-contest fame is now planning to offer a new prize '$5 million to $20 million for anyone who can decode the DNA of 100 or more people in a matter of weeks'. This gives one an idea of how important the once academic questions of biology have become relevant to the general masses. What would the next prize be . . . protein folding?
These are not the millions I was talkin about. These are just the tip of an ice-berg. By putting together 20 amino acids together biology has produced a plethora of protein structures and this was pretty much to be expected given the diversity that surrouds us. But even these different proteins are made of a large but limited number of motifs. Once all these motifs are charaterised one would truly have the nuts and bolts out of which actual functional proteins - molecular machines could be made. This would open up an entire industry of protein engineering. A good part of which may focus on chemically tweaking the existing proteins and a maybe a smaller group may focus on inventing entirely novel proteins. The possibilities seem to be endless. People are already using biological macromolecules in household products - e.g. Subtilisin a proteolytic enzyme is used in these.
I can go on and on . . . got the point ?
Wednesday, January 11, 2006
IM Gaim
 Of men and women and there IMs...
Of men and women and there IMs...Why am I rambling about some such topic? Is it really unlike of me to jabber on non-tech topics... :-? ewwwwwwwww lets explore the not-so-tech side...of people's IM-habits, IM-eccentricities, IM-egos, IM-hyphenated things.....
Lets start with a Sunday morning newspaper MCQ
Quantitating ur IQ: IM Quotient
1. Buddies Count?
a. Who counts...c'mmon
b. ummmmm
c. The messenger length/breadth is higly distorted (lengthwise)
2. Buddy Online/Offline ratio?
a. very very less than 1
b. approx 1
c. much greater than 1
3. Who's the 'initiator'?
a. me...i am a very good guy/gal ya know
b. depends
c. how do u do that? every time I login IM screens mysteriously pop up everywhere
4. Ur IM input/output ratio:
a. I am gr8 at typing ya know...my im windows are splattered with my say
b. Neva thot bout that really
c. **shrugs**
5. Away message
a. Who's away? me ...r u kiddin me
b. Sorry, I ran out for a bit! (classic Gaim)
c. i did'nt know ya miss mi so mach laddie
Ok lets see how U scored:
Mostly As: Its CyberSpace in case u did'nt notice...u can do without being urself..."Wake Up"
Mostly Bs: Well U r managing it well fella.
Mostly Cs: Ok...you dont have to be truthful evertime
After that taxonomy of IM creatures lets chk out something more about the IM-culture.
Dogs and bitches...
Hyenas, spiders, bees, dogs and other of those NAT GEO creatures. All these species have one common characteristic - the male must make the first move. Humans stick to their animal instincts in the IM land too. When a female collegue/ friend/ UFOD (undefined-female-object-of-desire)/...sorts... come online it is the moral responsibility of the male to 'initiate'. Female initiations are kinda rare cosmological events and need careful investigation.
Cant we just leave behind all our inhibitions when we are atleast IMing?
Should I/ Shouldn't I moments...
Okay so u login to ur account. U see this list of online buddies. U r not exactly in a mood to chit-chat. Okay with some of them, but not others. It is considered to be rude if you r online and donot IM with other 'available' onlines. So wat u end up doin is waste some bandwidth with some non-sense for 2 mins. Hang up with regular 'tc's and 'bbye's and ...the sorts. Why do all that? If u wanna chat plz initiate, if they want let them do it.
Bloated Egos...
Giving out ur IM id to ur boss is a very BAD idea. Bosses expect u to initiate, to do most of the chattering, entertain them...Besides the bosses, some other people around you who are'nt exactly ur bosses but create such an impression are like pseudo-bosses. These are the domineering sorta people. They have a kind of air to themselves. They will reply least frequently, with minimalist use of words, with huge latencies.
The END Problem...
Okay...this is not the end problem of molecular biology (d-uhh DNA replication)....this is a classic IM end problem. Let me put it straight - "How does one end an IM conversation?" People have this habit of trying to end the convo by these one liners:
tc
gnite
bbye
cya
ciao
chao
c u around
lets part
over n out
. . .
But no matter what u type...the person on the other side has to come up with some reply so that he-may-have-the-last-say.
The Solution: Well...it seems the problem has been solved by biology. When u wanna end a convo just type UGA (or UAA or UAG). WHY? Well go n revise ur high school biology. UAA, UGA n UAG are stop codons. What bout the non-science people U ask? In case U did'nt notice this is basically a tech blog and the intended audience is obviously other techies/nerds/geeks. What bout non-bio-science-people U ask? Well... if U r a non-bio-science-fella ; I hate to be the 1 to break it to ya fella but "U dont fit into the l33t group, aadeos" UGA.
more, comming soon...
Sunday, December 04, 2005
Appreciating Randomness...E.coli Interupted
Presenting a classic problem in genetics...a great way to introduce oneself to Mr. Poisson (of the distribution fame)...the joy of modelling...
[/mycrap]
Problem: The transfer of DNA from Hfr begins at oriT. The gene closest to oriT on the donor chromosome enters the recepient cell, followed by successive genes on the donor chromosome. The frequency of a particular gene entering the recepient is a function of its distance from oriT. Moreover, this frequency is found to decrease "exponentially" with a gene's distance from oriT. Why is this decrease exponential (and not linear, say)?
The Solution:
Poisson says
"Let m = the mean number of successful events. The probability of k successful events is given by the formula -
P(k) = (e^-m * m^k) / k!"
The key to solving the given problem is to define the 'm' value. Let us define the m value as the 'number of breaks per unit length of the chromosome'. So if 'x' is the length of a given chromosome segment the number of breaks in x will equal 'mx'. Remember that mx is the average number of breaks in a population of chromosome segments of length x. That is some x-lengthers will have more that mx breaks and some less than mx and some may even have zero breaks.
Even a single break in a length x is sufficient to ensure that a marker at distance x from oriT is not transferred. In other words any non-zero number of breaks in x will prevent marker at x from being transferred.
Now consider a circular ds DNA E.coli chromosome. Consider markers located at distances x1, x2 and x3 respectively from oriT.
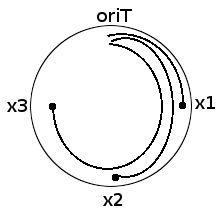
What are we distributing? Good question. We are distributing 'breaks' to x-length chromosome segment populations.
In case of x1-lengthers,
P(0) = Probability of having 0-break x1-lengthers = e^-mx1
P(1) = Probability of having 1-break x1-lengthers = mx1 * e^-mx1
P(2) = Probability of having 2-break x1-lengthers = (mx1^2 * e^-mx1) / 2
P(3) = Probability of having 3-break x1-lengthers = (mx1^3 * e^-mx1) / 6
.
.
.
Now comes the good part..
We are interested in whether a marker within distance x is transferred. It will be transferred if there are zero breaks in the length x. Hence we are interested in the fraction of those x-lengthers which have zero breaks. We are interested in the zero term P(0).
P(0) = e^-mx1
Similarly, fraction of markers at x2 transferred to the recepient,
P(0) = e^-mx2 and
fraction of markers at x3 transferred to the recepient,
P(0) = e^-mx3.
If we plot the values of P(0) for x1-lengthers, x2-lengthers and x3-lengthers versus the distance x1, x2, x3 then obvioulsy an exponentially decaying curve would be obtained. Multiplying the P(0) values of each by the total colonies scored will give the expected number of colonies recombinant for these markers, decaying exponentially with each's distance from oriT.
Thursday, November 17, 2005
Now Playing in Kopete...
Follow these simple steps to display the song currently playing in XMMS as your status in Kopete.
1. Create the following script - kopete-status.sh
#!/bin/sh
STATUS=" `current |awk '{ print substr($0,index($0,":")+2, index($0,"(")) }'` "
dcop kopete default setAway "$STATUS"
echo "Kopete Status: $STATUS"
2. Issue chmod u+x kopete-status.sh to make the script executable and then finally issue
watch -n 5 ./kopete-status.sh.
We r done. Watch your Kopete status being updated every 5 secs to the song being played in XMMS.
Note: U must have this package also: http://www.xmms.org/files/plugins/xmms-extra/xmms-extra-0.1.tar.gz
Friday, October 21, 2005
The Devil

I once heard someone important say "In biology, the devil lies in the detail".
Biology has since long become inter-disciplinary and is becoming even more so. Clearly, assuming that one has the analytical skills, more the number of facts one knows the better off he would be in biology.
It seems like until now I gave the devil a lot of free play.
Time to kill the devil...
Tuesday, September 13, 2005
Bottom Up
Thursday, September 01, 2005
Vacation at Binsar
Monday, August 22, 2005
Applause

I had blast!
Finally delivered the seminar today. Things went pretty smoothly.
I used the LCD projector for the first time. I loved every aspect of it. I did all the connections, wiring, setup...typical geekstuff.
There was no technical snag. But my own laser-pointer defied me :(
The words flowed out magically. I managed to convey atleast 50% of what I was saying. Partly becuse the 2nd half was kinda drab...perhaps too geeky for the un-enchanted-by-how-things-work.
I particularly liked my end-note:
"Our civilization has always been struggling to build tools which help us to measure time. First came the sand-clock, then the pendulum based ones and recently we have these hi-tech atomic clocks. But, ironically we all - the biologicals have had, right from the beginning an internal biological clock which keeps time faithfully. And now this clock [pause] this process has become aware of itself !"
Wednesday, August 17, 2005
A Simple Java Code
Code
import java.util.Vector;
public class MyCodons{
//globals
Vector codons = new Vector();
String[] bases = {"a","g","c","t"};
//fill the 'codons' vector with values
public MyCodons(){
for(int x=0; x<bases.length; x++){
for(int y=0; y<bases.length; y++){
for(int z=0; z<bases.length; z++){
codons.add(bases[x]+bases[y]+bases[z]);
}
}
}
}
//display the 64 codons
public void printCodons(){
System.out.println(codons.size()+" possible codons\n"+ codons);
}
//main method
public static void main(String[] args){
System.out.println("Generating codons...\n");
MyCodons obj = new MyCodons();
obj.printCodons();
}
}
Sunday, August 07, 2005
Molecular Logic: Biological Time Keeping

Going to give a seminar in the coming fortnight. We were given the freedom of choosing our topic. This topic hunt was something that I enjoyed.
I was reading a book 'Fly' - in which drosophila has been glorified as 'the' tool of molecular biology research. I stumbled upon a chapter on circadian clocks and got hooked. Drosophila apparently is the ideal model to study molecuar logic of circadian clocks.
The clock basically consists of three modules - the light input component, the oscillator and the output component. I am especially fascinated by biological light sensors, the first component. The idea that a clock can be implemented using transcriptional feed-back loops is really interesting, the second component. The role of the clock in organismal behaviour, the third component, immediately implies that this clock is not just another clock but an important control in organismal behaviour.
It seems that even a single aspect of this little insect, its clock, has a lot to offer. I should'nt have any problem gathering material for my upcoming 1 hr. speech.
Following is one of the interesting links I found on the web:
http://www.scienceden.com/mbiology/research/circadian
P.S.: The flash on this site is very helpful. As someone important has said "If a picture is worth a 1000 words what do you say about a moving picture !"
Thursday, August 04, 2005
H'ppy B'day 2 Me
Went to the Institute. Had no lectures today. Just had a stupid practical - Biuret Qualitative Analysis. Distributed Perk XL to buddies.
Came home. Mom had made mango and strawberry ice-cream. Enjoyed four scoops.
I really feel special.
Its good to be me.
Tom's Favorite Quotes and Mine Too
I was on a Google spree. Stumbled upon his really awesome collection of quotes.
Its a must read.
Tom's Favorite Quotes
P.S. Dont forget to listen to:
- fennyman's voice
- "Back off man, I am a scientist" - Ghostbusters. [ lmao ]
Wednesday, August 03, 2005
'Sound' Synthesis

I was thinking of an approach to do synthesis of sound - 'synthesis of voice' to be precise.
I have made few basic assumptions upon which the sound synthesis approach will be based. They are -
- speech is not continuous but it is made up of discreet elements.
- each individual has his own 'library of sound elements' from which s/he assembles speech.
- a library has one atrribute - number of elements (size).
- each element also has one attribute - the sound (note) which we hear.
- the libraries of any two individuals are of same size and contain the same types of notes.
- two analogous notes from any two different libraries differ only slightly so as to accomodate the fact that no two voices are same.
More to come ...


